
Do we predict mismatches that induce de novo DSA or do we predict the ones that do not?
Vadim Jucaud1, Lorita M. Rebellato2, Kimberly P. Briley2, Carl E. Haisch2, Scott A. Kendrick3, Heather Jones3, Kristel Mclawhorn3, David Leeser2, Matthew J. Everly1.
1Terasaki Research Institute, Los Angeles, CA, United States; 2East Carolina University, Greenville, NC, United States; 3Eastern Nephrology Associates, Greenville, NC, United States
The development of de novo donor-specific anti-HLA antibodies (dnDSA) remains a major risk factor for poor long-term kidney allograft outcomes, yet the prediction of dnDSA development has not been fully elucidated. Several molecular features (MoFs) of HLA mismatches (MMs) are currently used to predict the development of dnDSA. The purpose of this study is to evaluate the independent and concomitant impact of several MoFs of HLA MMs on the development of dnDSA, and the potential of machine learning techniques to predict dnDSA development.
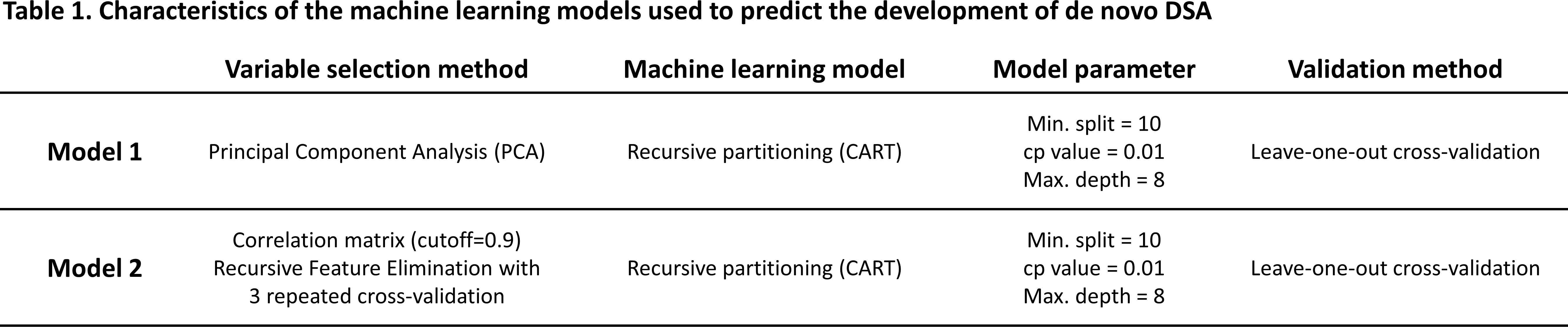
We retrospectively studied 236 HLA-mismatched kidney recipients transplanted between 2011 and 2016. All recipients received calcineurin inhibitors, did not have preformed DSA, were HLA-typed by NGS for HLA-A, -B, -C, -DRB1, -DRB345, -DQB1, -DPB1, and were screened for dnDSA using Single Antigen Beads with a median follow-up of 2.26 years. We used E3, an HLA immunogenicity software, to generate 55 MoFs of HLA MMs related to physiochemical differences, HLA antibody epitopes and HLA T cell epitopes, including the following commonly used MoFs: amino acid mismatch load (AML); electrostatic charge difference (ECD); hydrophobic difference (HD); mismatched HLA antibody epitope load (AbEpiL); and PIRCHE-II. To predict the development of dnDSA against specific HLA MMs, we used a recursive partitioning model with different variable selection methods, and a leave-one-out cross-validation (Table 1).
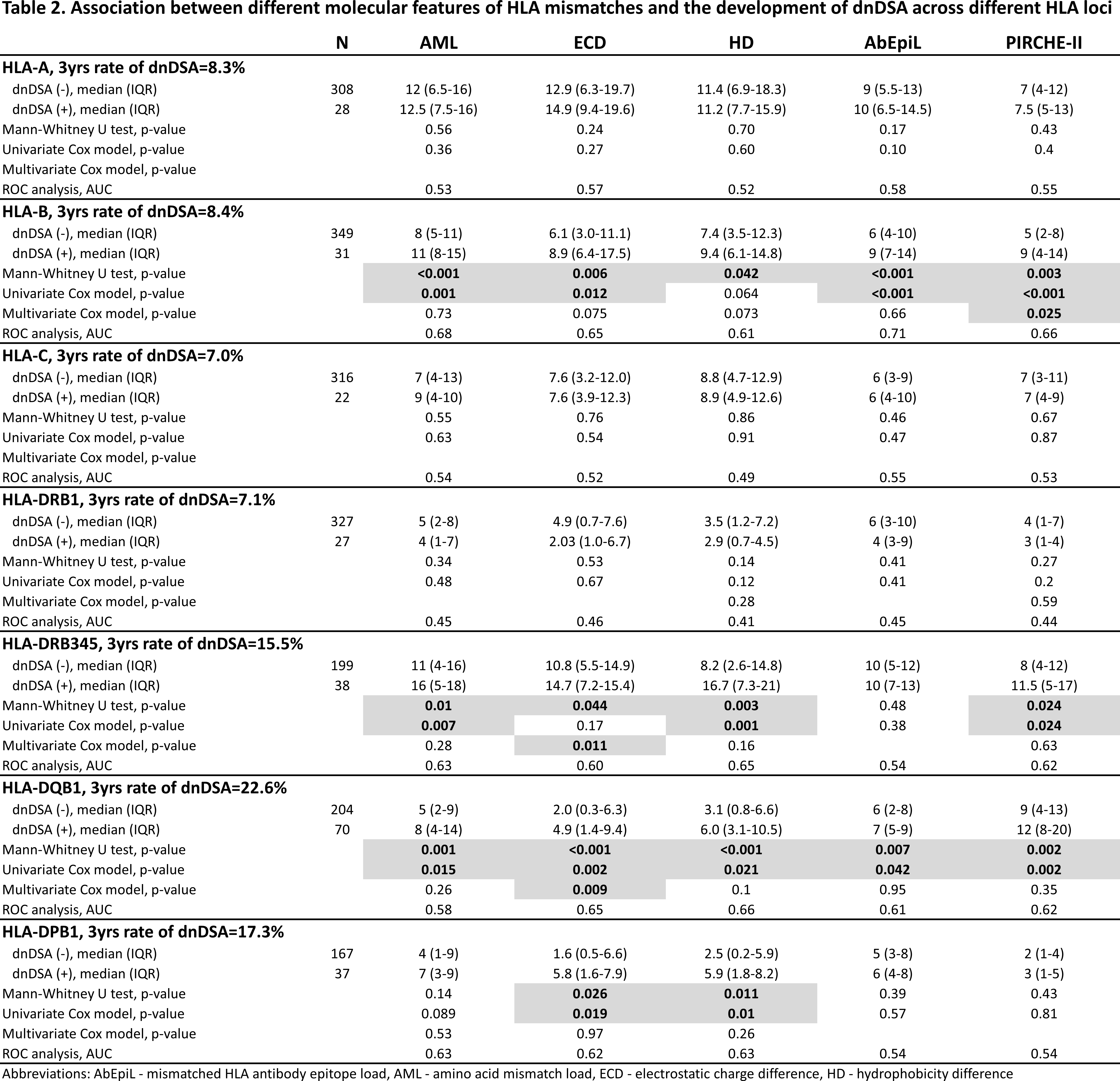
Among the 236 recipients, we observed a total of 2123 HLA MMs, of which 253 induced a dnDSA. The 3 year cumulative event rate of dnDSA development against the different HLA loci was statistically different (p<0.001) (Table 2). Overall, none of the common MoFs were strong predictors of dnDSA development (AUC range across HLA loci and MoFs: 0.41-0.71), although some were significantly associated with the development of dnDSA in both univariate and multivariate analyses for HLA-B, -DRB345, -DQB1, and -DPB1 only (Table 2
In both machine learning models, the prediction of dnDSA development was above 80% for HLA-A, -B, -C, and -DRB1 MMs, and below 80% for HLA-DRB345, -DQB1, and -DPB1. For model 1 and 2 respectively, the range of overall sensitivity was 52-73% and 22-55% and overall specificity was 96-99% and 96-99% across HLA loci (Table 3).
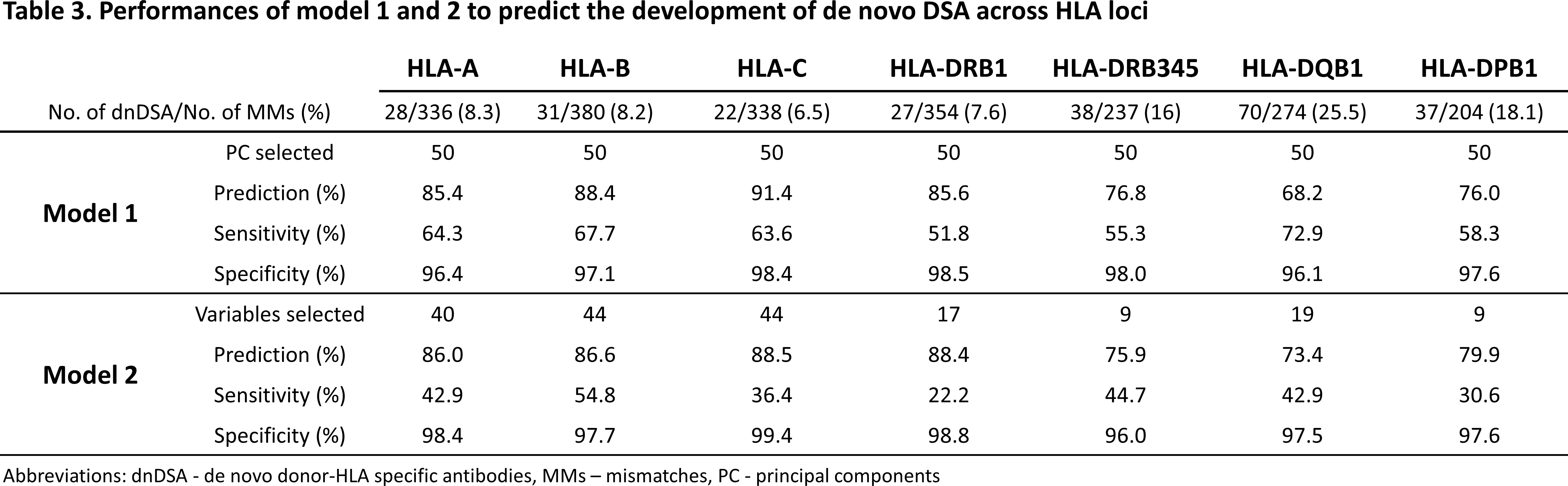
Last, the apparent low sensitivity of both models suggests that the correct identification of immunogenic HLA MMs remains poor, in spite of model 1’s sensitivity being higher than model 2.
None of the common MoFs of HLA MMs were strong predictors of dnDSA development. Yet, machine learning approaches suggest that development of dnDSA can be predicted accurately, but the prediction only reveals which HLA MMs will not induce a dnDSA rather than which HLA MM will induce a dnDSA. Nonetheless, it remains important – from a clinical perspective – to know if certain HLA MMs have a low risk of inducing a dnDSA.
The Terasaki Family Foundation.