The tissue Common Response Module (tCRM) score provides a quantitative objective biopsy score to assess the severity of pancreas transplant rejection
Yvonne Kelly1, Arya Zarinsefat1, Tara Sigdel1, Raphael Meier1, Minnie Sarwal1, Peter Stock1, Zoltan Laszik2.
1Surgery, University of California, San Francisco, San Francisco, CA, United States; 2Pathology, University of California, San Francisco, San Francisco, CA, United States
Introduction: Several studies have used gene expression profiling to identify key immunologic and inflammatory markers in the blood and tissue of solid-organ transplant recipients experiencing rejection (AR), but none of these have included samples from pancreas transplant (PT) recipients. Gene expression analysis of public transcriptional data from kidney, heart, liver, and lung transplant rejection led to the tissue Common Response Module (tCRM) score based on 11 genes (BASP1, ISG20, PSMB9, RUNX3, TAP1, NKG7, LCK, INPP5D, CXCL9, CD6, CXCL10; Khatri et al, 2013) which can diagnose the severity of tissue rejection, but this has not been validated in PT. In this study, we aimed to evaluate the locked tCRM score and identify additional markers of PT-AR.
Materials and Methods: We performed a retrospective study applying gene expression analysis to RNA isolated from formalin-fixed paraffin-embedded (FFPE) tissue from 51 pancreas biopsies, grouped by histopathologic grade of acute cellular rejection (Grade 1 (n=14), Grade 2 (n=14), and Grade 3 (n=9)) versus normal (n=14). Gene expression analysis was performed using the Nanostring Cancer Immune v1.1 oligonucleotide set, which includes 40 set housekeeping (HK) genes. Appropriate HK genes were selected, data was log transformed and normalized. Differential gene expression (DE) analysis involved a negative binomial model, unsupervised hierarchical clustering, and calculation of the tCRM score with removal of Nanostring designated HK genes confounded by rejection. P-value was considered significant at <0.05 (Benjamini-Yekutieli, Mann-Whitney, Graph-Pad PRISM). Pathway analysis (PA) was performed to evaluate biological significance.
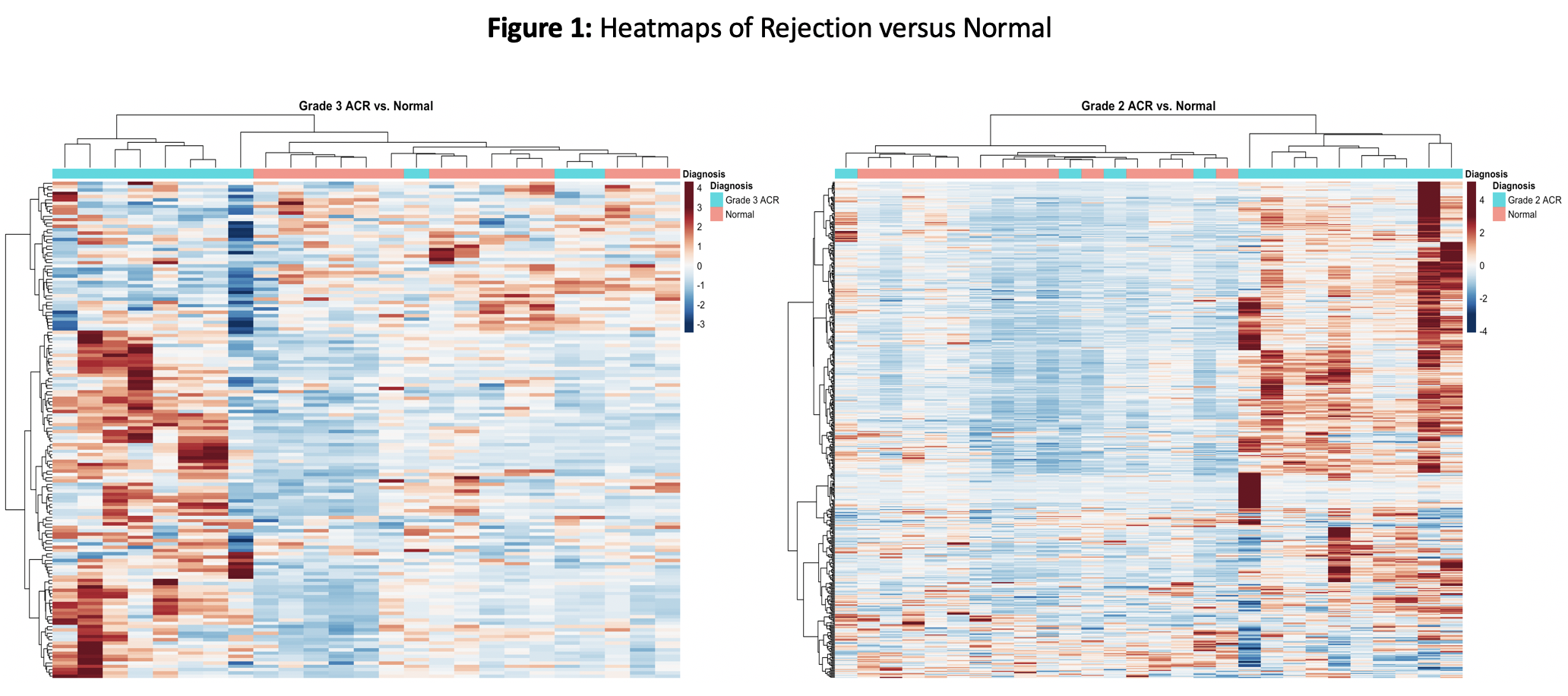
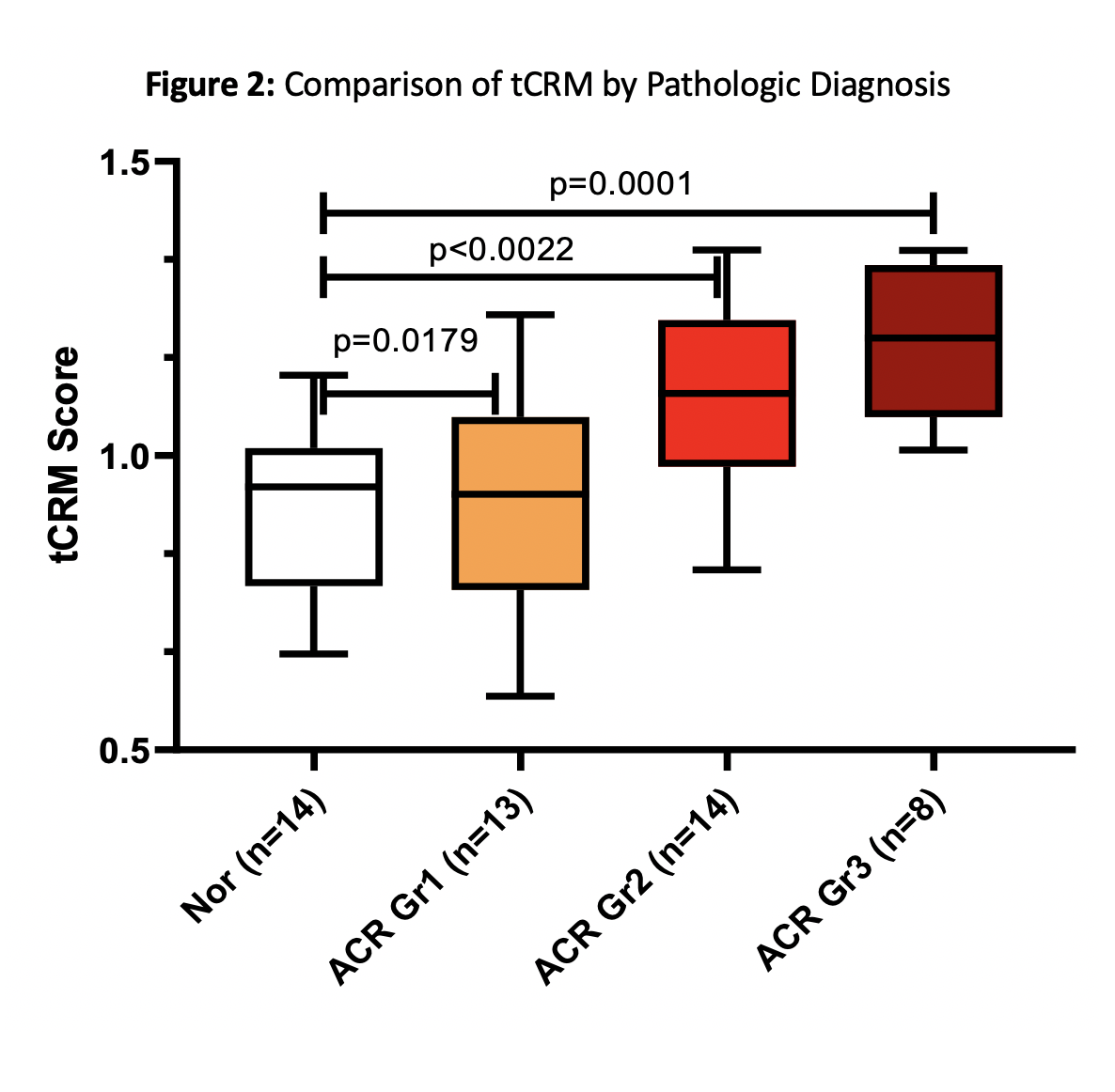
Results: Some Nanostring designated HK genes showed significant differences with AR and hence a custom set of HK genes was used for normalization of log transformed data. Significant transcriptional differences were seen with higher grades of rejection in PT (Fig.1). The tCRM score, run on a sub-component of 11 genes, captures the same message with a scaled score that increases across rejection grades (Fig.2). Of the 22 genes differentially expressed in Grade 3, 18 were also significant in Grade 2. Specifically, within the 11 tCRM genes diagnostic of rejection in other solid organ transplants, 9 of 11 were significant for PT rejection, with the last two tCRM gene (CXCL9 and CXCL10) approaching significance at p=0.055 and 0.08 respectively.
Conclusion: This study shows that a common hub of immune mediated genes, highly significant in transplant rejection (tCRM), and not confounded by tissue source (kidney, heart, liver and lung), were also highly informative for diagnosis and quantification of AR injury in PT. The bulk of the AR signal in PT, similar to other solid organs, is attributable to infiltration of activated leukocytes in the tissue, with background tissue injury specific transcriptional changes. The clinical and molecular drivers for Grade 1 PT rejection deserve additional analysis.
There are no comments yet...